Multi-Omics in Rare Disease Studies: Case Examples
Integrating genomics, transcriptomics, proteomics and metabolomics improves rare disease diagnosis and uncovers mechanisms guiding treatment.

Multi-omics is transforming how rare diseases are diagnosed and treated by combining multiple biological data layers - genomics, transcriptomics, proteomics, and metabolomics. This approach goes beyond identifying genetic mutations to uncover how these mutations disrupt cellular processes, enabling researchers to better understand disease mechanisms and identify potential treatments.
Key takeaways:
- Genomics identifies DNA mutations but often misses functional impacts.
- Transcriptomics tracks gene activity, revealing which genes are turned on or off.
- Proteomics measures protein levels to understand cellular functions.
- Metabolomics examines biochemical changes, offering insights into disrupted pathways.
Case studies demonstrate how multi-omics has:
- Improved diagnostic rates for rare metabolic diseases like methylmalonic aciduria (MMA).
- Identified genetic risks for conditions like sinusoidal obstruction syndrome (SOS) using integrated datasets.
- Solved undiagnosed cases by linking genetic variants to functional evidence.
- Explained phenotypic variability in patients with the same genetic mutation, aiding personalized treatment.
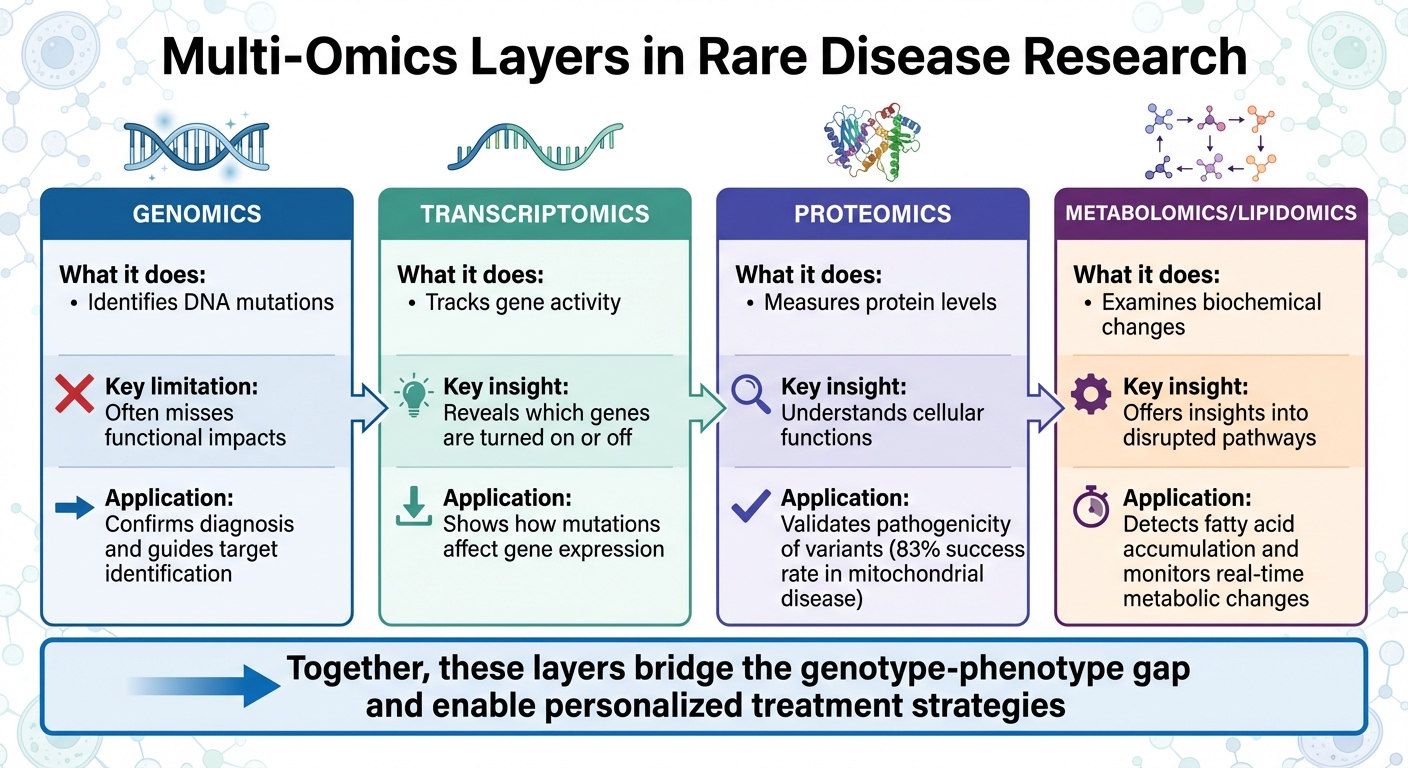
Multi-Omics Layers in Rare Disease Research: Genomics to Metabolomics
WEBINAR "Multiomics and Biomarkers in Rare Diseases Supporting Development of Orphan Drugs"
Case Study 1: Finding Genetic Risks in Sinusoidal Obstruction Syndrome
Sinusoidal obstruction syndrome (SOS) is a rare but severe complication that can arise after hematopoietic stem cell transplantation (HSCT). In its most extreme form, SOS can lead to multiple organ failure, with a mortality rate exceeding 80%. One of the biggest challenges in managing SOS is identifying patients at risk before symptoms develop. In April 2023, researchers from the University of Geneva and CHU Sainte-Justine tackled this issue by combining two omics datasets to uncover genetic risk factors.
Multi-Omics Pipeline for SOS Research
Led by Nicolas Waespe and Marc Ansari, the research team designed a pipeline that combined laboratory experiments with genetic data from patients. They began by treating six lymphoblastoid cell lines with busulfan, a chemotherapy drug known to trigger SOS. Using RNA sequencing, they identified 3,093 genes with altered activity: 1,708 were upregulated, and 1,385 were downregulated.
Next, the team conducted whole-exome sequencing (WES) on 87 pediatric HSCT patients, 12 of whom developed SOS, while the remaining 75 did not. The innovative aspect of their approach was integrating the transcriptomic and genetic data using a weighted Z-method. This method combined statistical signals from both datasets to enhance their ability to detect meaningful associations. As the researchers noted:
"This novel data analysis pipeline integrates two independent omics datasets and increases statistical power for identifying genotype-phenotype associations."
The results were striking. While traditional WES analysis identified only 4 SOS-associated genes, the integrated approach uncovered 35 genes linked to the condition. This marked a significant step forward in understanding the genetic underpinnings of SOS.
Findings and Clinical Impact
The 35 genes identified were closely tied to biological processes such as the p53 signaling pathway, apoptosis, and inflammation. Notable genes included:
- NACAP1: Linked to endothelial dysfunction.
- PF4V1: Associated with platelet disorders and clot formation.
- CYP7B1: Related to neonatal liver disease.
These findings shed light on the molecular mechanisms of SOS, particularly how busulfan damages blood vessel linings and triggers clotting. Building on this, the team developed a polygenic risk score (PRS) based on 209 genetic variants within the 35 identified genes. This score effectively distinguished SOS patients from controls, with a highly significant p-value of 8.506 × 10⁻⁷.
Clinically, this PRS has the potential to transform SOS management. By screening HSCT patients before transplantation, clinicians could identify those at high risk and provide early interventions. For example, high-risk patients could receive defibrotide, an anticoagulant and anti-inflammatory drug, as a preventative measure. Defibrotide is most effective when administered early, offering a chance to prevent SOS and save lives.
This case highlights how integrating multi-omics data can bridge the gap between genetic discoveries and practical clinical applications, offering new hope for improving outcomes in rare and life-threatening conditions like SOS.
Case Study 2: Multi-Omics in Pediatric Rare Disease Diagnosis
When it comes to diagnosing rare diseases in children, standard genetic tests often leave many questions unanswered. Whole-exome sequencing (WES) and whole-genome sequencing (WGS) can only identify the root cause in 31%–47% of cases, leaving families in diagnostic limbo for an average of 4.8 years. Multi-omics approaches are beginning to change this by uncovering genetic abnormalities that traditional methods simply miss.
Optical Genome Mapping for Genetic Variants
In May 2021, a team from the Undiagnosed Diseases Network (UDN), including Heidi Cope and Dr. Vandana Shashi, encountered a challenging case: a 4-year-old boy with severe epilepsy and developmental delays. Despite four years of testing - including chromosomal microarray analysis, a 38-gene epilepsy panel, and both exome and genome sequencing - no diagnosis was found.
The researchers turned to Optical Genome Mapping (OGM), using the Bionano Saphyr System. Unlike next-generation sequencing (NGS), OGM can detect structural variations that might otherwise go unnoticed. In this case, OGM identified a mosaic deletion and inversion spanning 90 kb on the X chromosome. This disrupted the CDKL5 gene and was present in 23.6% of the analyzed molecules - a level of mosaicism undetectable by standard methods.
This discovery provided the answer: X-linked dominant early infantile epileptic encephalopathy-2. As the research team explained:
"OGM affords an effective technology for the detection of SVs, especially those that are mosaic, since these remain difficult to detect with current NGS technologies and with conventional chromosomal microarrays."
This breakthrough with OGM paved the way for incorporating proteomics to tackle other diagnostic challenges, such as mitochondrial disorders.
Proteomics in Mitochondrial Disorders
While genomics is powerful, proteomics adds another layer of clarity, especially in mitochondrial diseases. Even with advanced genetic testing, over half of suspected mitochondrial disease cases remain unresolved. In July 2023, researchers from Monash University, led by Sumudu S.C. Amarasekera and David A. Stroud, used quantitative proteomics to address this issue in three pediatric patients with Leigh syndrome spectrum, a severe mitochondrial disorder.
For these "exome-unsolved" cases, the team applied tandem mass tag proteomics to fibroblast cell lines, analyzing around 8,000 proteins per sample. This revealed a clear reduction in large mitoribosomal subunits, suggesting impaired mitochondrial protein synthesis.
Using this protein signature, they revisited the patients' genomic data and identified biallelic variants in MRPL39, including a deep intronic mutation that introduced a cryptic exon. This type of mutation is typically missed by standard exome sequencing. The researchers noted:
"Our study highlights the utility of quantitative proteomics in detecting protein signatures and in characterizing gene-disease associations in exome-unsolved patients."
Elsewhere, in Spain, researchers have used mass spectrometry-based proteomics to study rare inherited metabolic diseases. Their work has identified biomarkers for conditions like methylmalonic aciduria, Fabry disease, and various mucopolysaccharidoses - cases where genetic tests alone often fall short. Together, these examples show how multi-omics approaches can solve diagnostic puzzles that traditional methods cannot.
Case Study 3: Solving Undiagnosed Cases with Multi-Omics Cohorts
For many families, traditional genetic testing often leaves questions unanswered. Even with advanced methods like whole-exome or whole-genome sequencing, only 35–70% of cases result in a molecular diagnosis. One of the biggest hurdles is confirming whether a genetic variant is truly disease-causing. Multi-omics cohort studies tackle this challenge by combining data from multiple cases to validate diagnostic signals and provide the functional evidence needed to solve these complex cases.
Matched Omics for Mitochondrial Diseases
In July 2023, researchers at Monash University faced a tough diagnostic puzzle involving three unrelated children with severe multisystem disease suspected to stem from mitochondrial disorders. Exome sequencing didn’t provide answers. The research team, led by Sumudu S.C. Amarasekera and David A. Stroud, turned to quantitative proteomics, analyzing fibroblast samples from the patients. This approach uncovered a specific reduction in large mitoribosomal subunits - proteins critical for mitochondrial function - suggesting a defect in mitochondrial protein synthesis.
With this clue, the team re-examined genomic data and performed whole-genome sequencing, which revealed a shared deep intronic variant in the MRPL39 gene. This mutation, missed by exome sequencing, was found to create a cryptic exon that disrupted normal protein production. RNA sequencing confirmed the finding, finally delivering a diagnosis for these families. Inspired by this success, larger multi-omics cohort studies have since validated this diagnostic framework, showing how these integrated methods can address such challenging cases.
Broad Rare Disease Cohorts and Outcomes
The success of multi-omics approaches isn’t limited to mitochondrial disorders. In May 2025, the Victorian Clinical Genetics Services demonstrated how ultra-rapid proteomics could complement acute-care genomics for critically ill infants. One case, labeled UDP8, involved a newborn suspected of having a mitochondrial disease. Rapid trio whole-genome sequencing identified variants in the NDUFS8 gene, but functional validation was still needed. Using proteomics, researchers analyzed blood cells in just 54 hours, identifying a significant reduction in NDUFS8 protein and an isolated complex I defect. This functional evidence confirmed the diagnosis.
Across broader cohorts, proteomics has consistently outperformed traditional methods. For example, proteomics testing supported the pathogenicity of variants in 83% of individuals with confirmed mitochondrial disease, surpassing the results of clinical respiratory chain enzymology. As Associate Professor David A. Stroud from Monash University explained:
"This study provides a framework to support the integration of a single untargeted proteomics test into routine diagnostic practice for the diagnosis of mitochondrial and potentially other rare genetic disorders in clinically actionable timelines."
This shift from long diagnostic journeys to a rapid turnaround of about 54 hours represents a major leap forward. Multi-omics cohort studies are proving to be game-changers, delivering the functional evidence needed to diagnose rare diseases with speed and precision, especially for critically ill patients.
Case Study 4: Adding Proteomics, Metabolomics, and Lipidomics to Genomics
Understanding Phenotypic Variability
Genomic sequencing can identify mutations, but it often leaves a lingering question: why do individuals with the same genetic variant experience different symptoms? This variability in phenotypes can be perplexing for both clinicians and families. The key to this mystery lies beyond the genome itself - in the dynamic layers of biology such as proteins, metabolites, and lipids, which evolve in response to factors like environment, aging, and lifestyle.
In December 2021, a team led by Ling Hao at The George Washington University took on this challenge. They studied fibroblasts from patients with MELAS (mitochondrial encephalomyopathy, lactic acidosis, stroke-like episodes) who carried the ultra-rare pathogenic variant m.14453G>A. By integrating proteomics and metabolomics, they uncovered a significant deficiency in mitochondrial complex I and observed downregulation in arginine biosynthesis. These molecular insights explained the hallmark symptom of stroke-like episodes and supported the clinical practice of treating these patients with arginine infusion.
This approach addresses what researchers call the "genotype-phenotype gap." While DNA sequencing might identify a variant of unknown significance, proteomics can reveal whether the mutation disrupts protein production, and metabolomics can highlight cellular chemical imbalances that genomics alone might miss. Dr. Pankaj Agrawal, Chief of the Division of Neonatology at the University of Miami Miller School of Medicine, emphasized the importance of these technologies:
By bringing these other technologies into the mix, we can improve our ability to diagnose rare diseases and get answers for these families.
This level of detailed pathway analysis not only aids in understanding the disease but also paves the way for identifying actionable therapeutic targets.
Pathway Analysis and Disease Mechanisms
When researchers combine multiple layers of "omics" data, they can trace how information flows from DNA to RNA to proteins and metabolites, offering a clearer picture of how molecular processes contribute to disease. This type of pathway analysis often reveals therapeutic targets that would remain hidden if only genomics were used.
The MELAS study is a prime example. By analyzing proteins and metabolites together, the researchers identified a blockage in the argininosuccinate synthase enzyme within the arginine biosynthesis pathway. This discovery provided molecular evidence to back an existing treatment, turning a suspected mechanism into a validated therapeutic approach.
The potential of multi-omics extends beyond mitochondrial disorders. For instance, research into Miller-Dieker Syndrome highlighted repressed EIF2 signaling and reduced phosphorylation of mTOR pathway regulators, which are tied to defects in protein translation. Other studies have linked rare disease symptoms to disruptions in the methionine cycle and inositol lipid signaling, pointing to new opportunities for targeted treatments.
Lessons from Case Studies and RareLabs' Translational Applications
What We Learned from Multi-Omics Case Studies
From the case studies discussed earlier, one thing becomes clear: understanding rare diseases requires integrating multiple layers of omics data. Genomics can pinpoint mutations, but it's the combination with metabolomics and lipidomics that uncovers the disrupted biochemical pathways driving disease variability. By layering these approaches, researchers can connect genetic mutations to the biochemical changes that fuel disease progression.
RareLabs' Personalized Drug Discovery Programs
RareLabs has transformed these insights into a patient-focused approach for drug development, leveraging multi-omics integration. The process begins with a skin biopsy to generate patient-specific induced pluripotent stem cells (iPSCs), which act as personalized disease models. These cells, reflecting the patient’s unique genetic profile, allow researchers to directly test therapy effectiveness.
Therapeutic screening covers a wide range of approaches, including small molecules, antisense oligonucleotides, and gene replacement therapies, all tested on patient-derived cells. This personalized model ensures that potential therapies are validated using the patient’s own biological material. A compelling example comes from a June 2024 study by Apteeus and CHU Lille researchers. They used a bedside-to-bench-to-bedside approach for a patient with ACOX1 deficiency. By screening a library of repurposed drugs on the patient’s skin fibroblasts with mass spectrometry, they identified niclosamide, which successfully reduced toxic very long chain fatty acid (VLCFA) levels. The patient benefited from this treatment for over six years.
"We implemented screening technologies and mass spectrometry analysis which enable the measurement of the effect of small molecules directly on the patient's cells. This approach is particularly well-adapted to such an inherited metabolic disorder."
– Camille Moreau et al., Researchers
RareLabs also prioritizes transparency by keeping families informed with plain-language updates and offering direct access to its scientific team throughout the drug discovery process.
How Multi-Omics Strengths Align with RareLabs' Offerings
The table below outlines how various omics layers contribute to rare disease research and how RareLabs incorporates these insights into their therapeutic development:
| Omics Layer | Key Advantages | Application at RareLabs |
|---|---|---|
| Genomics | Identifies primary genetic variants and disease risks | Confirms diagnosis and guides target identification for screening |
| Lipidomics | Detects accumulation of specific fatty acids (e.g., VLCFA in peroxisomal disorders) | Validates drug efficacy through mass spectrometry on patient cells |
| Metabolomics | Monitors real-time metabolic changes and pathway disruptions | Measures biochemical impact during high-throughput drug screening |
| Patient-Derived Cells | Provides a personalized "patient-in-a-dish" platform | Enables testing of therapeutic candidates on an individual's own biology |
This approach not only simplifies target identification but also enables precise validation, especially for ultra-rare diseases. RareLabs uses the strengths of each omics layer to bridge the gap between molecular findings and personalized treatments. For conditions where traditional clinical trials are not feasible, this individualized strategy acts as a functional "n-of-1" trial, providing early evidence of a therapy’s effectiveness before long-term use.
Conclusion
Case studies reveal how multi-omics extends beyond genomics by incorporating transcriptomics, proteomics, metabolomics, and lipidomics. This combination delivers the functional insights needed for accurate diagnoses and targeted treatments.
Dr. Pankaj Agrawal, Chief of the Division of Neonatology at Miller School of Medicine, underscores the importance of this approach:
It's important that we leave no stones unturned for these patients. If the genomic test is inconclusive, we have other technologies available to help get answers.
One standout example is the June 2024 ACOX1 deficiency case, where a skin biopsy became the cornerstone for personalized drug screening. This led to over six years of effective treatment with niclosamide. By using patient cells as models for drug testing, researchers can evaluate therapies directly against an individual’s unique biology.
RareLabs takes this concept further, transforming skin biopsies into patient-derived iPSCs for personalized drug screening. By integrating mass spectrometry and metabolomics, they measure biochemical markers to confirm whether treatments address the root metabolic issues. This approach redefines rare disease therapeutics by blending multi-omics innovation with patient-specific modeling.
For ultra-rare conditions, where traditional clinical trials are often unfeasible, this method offers a practical alternative. It allows for individualized solutions tailored to inherited metabolic disorders, benefiting patients and advancing science and medicine. RareLabs exemplifies this philosophy, turning multi-omics research into treatments customized to each patient’s genetic and biochemical profile.
The future of rare disease research lies in more than just gathering data - it's about transforming that data into actionable models that fuel therapeutic breakthroughs. These case studies demonstrate the potential of this approach, and platforms like RareLabs are bringing these advancements to patients and their families.
FAQs
When should multi-omics be used after WES or WGS?
Multi-omics is often suggested as a follow-up to whole-exome sequencing (WES) or whole-genome sequencing (WGS) when more detailed insights are required to unravel complex genetic conditions. By combining data from transcriptomics, proteomics, and metabolomics, it provides a broader molecular perspective. This approach can be especially helpful when WES or WGS results are unclear or don't entirely explain a patient’s phenotype, improving diagnostic precision for rare diseases.
How can proteomics or metabolomics prove a variant is pathogenic?
Proteomics and metabolomics play a key role in showing whether a variant is pathogenic by highlighting major disruptions in protein, metabolite, or cytokine profiles associated with the disease. These disruptions serve as functional evidence, demonstrating how the variant impacts biological pathways. For instance, studies on mitochondrial diseases and primary immune deficiencies have successfully used this approach to uncover such connections.
What does a patient-derived cell model enable at RareLabs?
Patient-derived cell models are often a cornerstone in personalized research. While the specific role they play at RareLabs isn't detailed, these models are generally used to delve deeper into rare genetic diseases. By studying cells that come directly from patients, researchers can gain insights into the unique characteristics of these conditions and work toward developing therapies that are tailored to individual needs.



