Study Insights: Multi-Modality Drug Discovery
Integrated omics plus AI improve rare-disease diagnosis and target ID, enabling personalized, combination therapies tested in patient-derived models.

Multi-modality drug discovery is changing how we understand and treat diseases, especially rare genetic disorders. By integrating multiple data types - like DNA, RNA, proteins, and metabolites - this approach provides a more complete view of disease mechanisms that single-modality methods often miss. Here are the key takeaways:
- Better Diagnostics: Combining whole-genome sequencing (WGS) with RNA sequencing and proteomics improves diagnostic success rates from 47% to 54% in rare diseases.
- Improved Drug Development: AI and deep learning tools like Graph Convolutional Networks (GCNs) are uncovering new therapeutic targets and pathways, such as NF1 in breast cancer and BACE1 in Alzheimer’s.
- Combination Therapies: Multi-modality approaches enable the design of effective combination treatments, such as pairing RNA-based therapies with small molecules for diseases like ALS and cancer.
- Patient-Centered Research: Using patient-derived models, researchers are developing personalized treatments faster and with higher precision.
This shift is supported by new tools like the updated Therapeutic Target Database (TTD) and regulatory changes encouraging personalized medicine. With multi-modality strategies, researchers can address complex diseases more effectively, offering hope for patients with limited options.
Multimodal AI in Medicine and Drug Discovery
Recent Advances in Multi-Omics Data Integration
Integrating multiple layers of biological data has always been a tricky puzzle, but recent breakthroughs in computational tools are making it more manageable. Deep learning frameworks are now capable of combining genomics, transcriptomics, and proteomics to uncover patterns that traditional methods often miss. These developments build on earlier discussions about multi-modality drug discovery, offering greater precision in identifying therapeutic targets.
Deep Learning Models for Data Integration
A key approach in modern data integration is intermediate fusion. This method processes each type of data independently before combining their core features. Models such as Graph Convolutional Networks (GCN), Graph Attention Networks (GAT), Variational Autoencoders (VAEs), and transformers excel at handling multi-omics data. They achieve this by preserving the structure of biological networks, filtering out noise, and prioritizing the most relevant evidence. Tools like MoRE-GNN go a step further by dynamically constructing relationship networks directly from the data, tailoring the analysis to each dataset's unique characteristics.
Examples of Multi-Omics Studies
The real-world impact of these deep learning models is best seen in specific studies:
- At Washington University, researchers developed MosGraphFlow, an AI model that identified key biomarkers and signaling interactions linked to Alzheimer's Disease progression.
- Zhanpeng Huang's team at Guangdong Pharmaceutical University introduced MOGOLA, a framework that combines GCN and GAT architectures with Omics-Linked Attention. This model, tested on breast cancer and Alzheimer's datasets (BRCA and ROSMAP), outperformed 11 other methods.
- MORGaN, a self-supervised model created by Queen Mary University of London and Recursion Pharmaceuticals, analyzed pan-cancer data from The Cancer Genome Atlas. It improved target identification by 9% and successfully prioritized drug targets associated with pharmaceutically important signaling pathways.
- La Ode Aman's group demonstrated how integrating five omics layers with protein network data using GCNs could predict cancer cell responses to Dabrafenib with an impressive R² of 0.96.
These examples highlight how deep learning is reshaping multi-omics research, offering tools that not only analyze data but also provide actionable insights for disease understanding and treatment development.
Therapeutic Targets and Pathway Insights
Key Targets Identified in Recent Studies
Recent advancements in multi-modality research are shedding light on molecular targets hidden within the complexity of multi-gene disorders. For instance, the multimodal transformer Precious1GPT has pinpointed APLNR and IL23R as significant regulators in age-related diseases. APLNR is involved in maintaining cardiovascular and metabolic health, while IL23R acts as a receptor for proinflammatory cytokines. These findings are especially relevant as global life expectancy is anticipated to climb to 77.2 years by 2050.
In the field of oncology, NF1 (Neurofibromin) has gained attention as a pivotal target in breast cancer. Mutations in NF1 lead to continuous activation of the Ras, PI3K/AKT, and MAPK signaling pathways. To counteract this, researchers have utilized antisense oligonucleotide (ASO)-mediated exon skipping to restore functional neurofibromin. Similarly, in Alzheimer’s disease, multi-task neural networks have highlighted BACE1 and GSK-3β as dual-target candidates to tackle both amyloid aggregation and neuroinflammation. Multi-omics Mendelian randomization has further validated several high-confidence targets for Alzheimer’s, including PILRA, GRN, ACE, TIMD3, and TREM2.
These breakthroughs naturally lead to a deeper understanding of how these targets interact within complex biological pathways.
Pathway-Level Discoveries
Integrating these targets into pathway-level analyses is transforming therapeutic strategies by enabling the design of combinatorial treatments. Central to many multi-gene disorders are the PI3K-Akt and Ras signaling pathways. In cancers with NF1 mutations, the absence of neurofibromin causes hyperactivation of these pathways. A 2026 study utilizing a multi-organoid platform identified the drug Paxalisib, which, when paired with ASO-mediated exon skipping, demonstrated a synergistic anticancer effect.
Pathway dysregulation is not limited to cancer. In sporadic ALS - responsible for about 90% of ALS cases - research has revealed disrupted RNA metabolism, nucleocytoplasmic transport, and DNA damage repair pathways. A March 2025 study involving motor neurons derived from 100 sporadic ALS patients tested a combination therapy. The regimen, which included Baricitinib, Memantine, and Riluzole, significantly improved motor neuron survival. This is a notable achievement, given that 97% of previously tested drugs have failed to prevent neurodegeneration in similar models.
For disorders linked to defective protein interactions, the 14-3-3γ-phosphotarget pathway offers promising insights. Mutations in YWHAG disrupt 14-3-3γ’s ability to bind with critical phosphorylated partners like TH and LRRK2, leading to pediatric encephalopathy. Over 50% of YWHAG-related cases can be traced to mutations in specific amino acids (Arg57, Arg132, and Tyr133). A high-throughput screening of 3,000 approved drugs identified Nafamostat mesylate as a strong candidate for drug repurposing.
In February 2026, researchers at the Broad Institute demonstrated the potential of the brain-selective mTORC1 activator NV-5138. During Phase II trials, this compound reversed cellular and molecular defects in cerebral organoids derived from lissencephaly patients.
"Clinically distinct genetic diseases converge on shared, druggable nodes" - Anna Greka, Mass General Brigham
Databases and Tools Supporting Drug Discovery
Therapeutic Target Database 2026 Updates

The Therapeutic Target Database (TTD) 2026 plays a key role in organizing drug discovery data. The latest update includes 306,247 target–disease associations across 2,912 targets, giving researchers a powerful tool to explore treatments for complex, multi-gene disorders.
One standout feature of this update is the addition of 10,506 perturbation profiles for 2,368 targets. These profiles capture how targets respond to knockout, overexpression, or chemical modulation - essential insights for predicting how potential drugs might perform in living systems. On top of that, the database now includes clinical profiles for 2,234 approved drugs, complete with FDA label information on adverse reactions and pharmacokinetics. This helps researchers identify safety concerns early in the drug discovery process.
Beyond TTD, other specialized databases are tackling unique challenges in drug research. For example, TRESOR (TWAS-relevant signature for orphan diseases) combines GWAS summary data with transcriptome-wide association studies to develop functional disease signatures. In 2025, TRESOR identified BECN1, FHL2, and CNOT7 as potential new targets for Multiple Endocrine Neoplasia (MEN), a rare condition that previously had only one known target. Meanwhile, the HCDT 2.0 database, updated in March 2025, offers a three-dimensional view of 1,284,353 curated interactions, including 1,224,774 drug-gene pairs and 11,770 drug-RNA mappings. These detailed profiles are laying the groundwork for innovative combination therapy strategies.
Applications to Combination Therapy Design
Modern databases are also proving invaluable for designing combination therapies by identifying co-occurring targets. For instance, TTD 2026's target co-occurrence insights reveal that the NET-SERT (norepinephrine transporter-serotonin transporter) pair is linked to 10 approved drugs, offering concrete evidence of this combination's therapeutic potential. Similarly, analyzing target pairs like JAK1-JAK2 or EGFR-HER2 helps researchers pinpoint "synergistic windows" where combining treatments can yield better results than targeting a single pathway.
Another tool, DeepTarget, integrates viability screens with CRISPR data to predict drug effects with remarkable precision. In 2025, it identified pyrimethamine (Daraprim) as a treatment for cancer by inhibiting mitochondrial oxidative phosphorylation - a finding later validated in U937 leukemia cells. DeepTarget also confirmed that EGFR serves as a secondary target for Ibrutinib in BTK-negative solid tumors, particularly in cells with the T790M mutation. These examples highlight how layered data - from genetic screens to clinical outcomes - can accelerate the journey from hypothesis to validated therapy, pushing drug discovery into new territory.
Implications for Personalized Drug Discovery
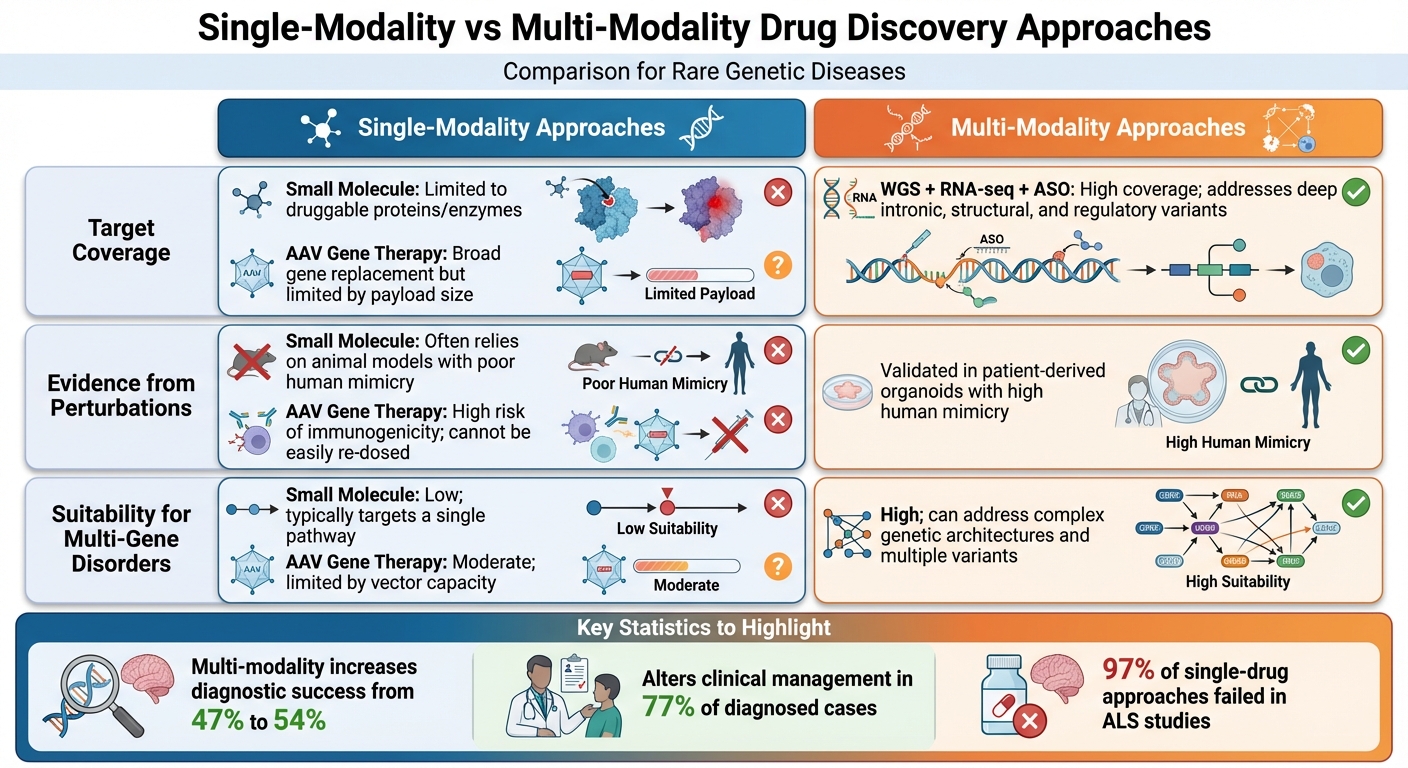
Single-Modality vs Multi-Modality Drug Discovery Approaches Comparison
Multi-Modality Strategies in Practice
The move toward multi-modality drug discovery is reshaping how researchers address rare genetic diseases. Rather than sticking to a single therapeutic route, teams are combining methods like small molecule screening, antisense oligonucleotides (ASOs), gene replacement, and combination therapies to tackle the intricate biology of multi-gene disorders. This layered strategy is already delivering tangible results using patient-derived specimens.
Take Duchenne muscular dystrophy (DMD) as an example. Researchers working with siblings affected by a deep intronic variant of DMD created cardiac organoids from patient cells and developed personalized ASOs. In just six weeks, this treatment restored dystrophin levels and normalized calcium flux in the cardiac organoids. A rapid induced pluripotent stem cell (iPSC) pipeline further demonstrated efficiency, achieving a 93% success rate in establishing patient-derived lines within three weeks.
Another study demonstrated the power of combining therapies. By integrating NF1 exon-skipping ASOs with Paxalisib, researchers observed synergistic anticancer effects. Meanwhile, a national study in Australia involving 290 families revealed that adding RNA sequencing and proteomics to standard whole-genome sequencing increased the diagnostic yield from 47% to 54%. This multi-omic approach also altered clinical management in 77% of diagnosed cases. In a separate study of 100 sporadic ALS patients, a triple-drug combination (baricitinib, memantine, riluzole) significantly improved motor neuron survival, whereas 97% of previously tested drugs had failed.
"Rare disease has pushed innovators to build new toolsets." – Jennifer Meade, President, Aldevron
These examples highlight the limitations of single-modality approaches, as explored in the following comparison.
Single vs. Multi-Modality Approaches
| Modality Type | Target Coverage | Evidence from Perturbations | Suitability for Multi-Gene Disorders |
|---|---|---|---|
| Single-Modality (Small Molecule) | Limited to druggable proteins/enzymes | Often relies on animal models with poor human mimicry | Low; typically targets a single pathway |
| Single-Modality (AAV Gene Therapy) | Broad gene replacement but limited by payload size | High risk of immunogenicity; cannot be easily re-dosed | Moderate; limited by vector capacity |
| Multi-Modality (WGS + RNA-seq + ASO) | High; covers deep intronic, structural, and regulatory variants | Validated in patient-derived organoids with high human mimicry | High; can address complex genetic architectures and multiple variants |
This table underscores why multi-modality approaches are better equipped to deal with complex genetic disorders.
Single-modality treatments often fall short when diseases involve multiple pathways or when compensatory feedback loops counteract the therapy. Multi-modality strategies overcome these hurdles by addressing disease biology from several angles at once. For instance, in Timothy Syndrome, researchers used an ASO-based "exon-switching" strategy to shift CACNA1C expression from the pathogenic exon 8A to exon 8. This approach corrected calcium defects in both organoids and rat models. This level of precision, validated in patient-derived tissues, addresses challenges that traditional drug development struggles to resolve.
RareLabs' Approach to Personalized Medicine
RareLabs builds on these multi-modality breakthroughs with a tailored three-step process that combines patient-derived models with cutting-edge therapeutic techniques. The journey begins with creating iPSCs from a simple blood draw, producing cellular models that replicate the patient's genetic profile. These cells are then corrected using CRISPR to generate healthy control lines, with karyotyping ensuring quality before therapeutic testing starts.
The second step focuses on personalized drug discovery, specifically designed for the patient’s genetic makeup. RareLabs examines options like small molecule screening, ASO therapy, gene replacement, and combination treatments to find the most effective strategy. This process is anything but standardized; every therapeutic plan is customized to the patient’s unique mutation and disease mechanism.
Throughout, families are kept informed with clear progress updates and milestone reports. They also have direct access to the science team. As Elisabeth Gardiner, PhD and CSO of Tevard Biosciences, explains:
"It's only rare until it's your loved one that has the rare disease, and then all of a sudden you realize that you're part of this club... desperate for answers."
RareLabs exemplifies the broader trend in rare disease research: moving away from generic methods toward personalized strategies that integrate various data sources and therapeutic options. The FDA’s February 2026 draft guidance for individualized therapies supports this shift, allowing the use of natural history data and small clinical studies for ultra-rare conditions. With gene therapies costing between $850,000 (Luxturna) and $3.5 million (Hemgenix), it’s crucial to get the treatment plan right on the first attempt. Multi-modality discovery, validated through patient-specific models, offers a promising way forward.
Conclusion
The field of multi-modality drug discovery is reshaping how rare genetic diseases are treated. Research demonstrates that combining whole-genome sequencing with RNA sequencing and proteomics increases diagnostic success rates from 47% to 54% while influencing clinical management in 77% of diagnosed cases. These integrated methods reveal therapeutic possibilities that single-modality approaches often overlook.
Patient-derived iPSC models have proven to be instrumental in identifying effective drug combinations for diseases like ALS. Additionally, multi-organoid platforms - integrating intestinal, liver, and kidney organoids - now allow for personalized drug profiling that evaluates treatment effects across entire biological systems, rather than focusing solely on isolated targets.
"Clinically distinct genetic diseases converge on shared, druggable nodes." – Jillian L. Shaw & Anna Greka, Broad Institute of MIT and Harvard
This convergence has sparked a shift in rare disease research. Advanced AI platforms like Madrigal are leading the way by integrating data from structural, pathway, viability, and transcriptomic analyses to predict clinical outcomes for over 21,842 compounds and 953 clinical endpoints. Regulatory support is also growing, with the FDA's February 2026 draft guidance for individualized therapies creating new pathways for personalized treatments, especially for conditions where traditional trials are impractical. These scientific and regulatory milestones are paving the way for more tailored and effective treatment strategies.
RareLabs exemplifies how these advancements are being applied in practice. By leveraging multi-modality techniques, RareLabs enhances precision in drug discovery. Their approach combines patient-derived iPSC models with tailored therapeutic strategies, such as small molecule screening, ASO therapy, and gene replacement. Testing multiple modalities in patient-specific models before choosing a treatment offers families a clearer path toward answers and solutions.
FAQs
What is multi-modality drug discovery?
Multi-modality drug discovery brings together different types of biomedical data and techniques - like imaging, genomics, and proteomics - to create new therapies. By combining these data sources with advanced AI tools, researchers can improve drug-target predictions, uncover new therapeutic targets, and fine-tune treatments. This approach is particularly valuable for tackling complex disorders involving multiple genes.
In the context of personalized medicine, this method plays a key role in tailoring treatments to individual patients. It supports the development of therapies such as small molecules, antisense oligonucleotides, and gene therapies, ensuring they meet the unique needs of each person.
How does adding RNA-seq and proteomics improve rare-disease diagnoses?
Adding RNA-seq and proteomics to rare-disease diagnoses provides a deeper understanding of how genes and proteins function within the body. These methods help uncover disease-causing variants that standard approaches might miss, leading to more accurate and quicker diagnoses. By offering a clearer picture, especially for intricate genetic conditions, these tools greatly improve the ability to pinpoint the root causes of rare diseases.
When do combination therapies work better than a single treatment?
Combination therapies have shown to be more effective than single treatments, especially when designed to address specific tumor mutations or molecular profiles. Recent research reveals that personalized multi-drug regimens can lead to better treatment outcomes. Additionally, studies show that responses to targeted inhibitors can vary across different models, emphasizing the importance of tailoring treatments. This method taps into precision medicine to tackle complex conditions with greater success.



